BAGGING, BOOSTING, and RANDOM FORESTS
1. Bagging and Random Forests
Recall that bagging is a random forest with m = p. So, here is just an example of a random forest.
> library(randomForest)
> rf = randomForest(mpg ~ .-name, data=Auto) # By default, m = p/3. But we can also choose our own m.
> rf # (m is the number of X-variables sampled at each node)
Call:
randomForest(formula = mpg ~ . - name, data = Auto, subset = Z)
Type of random forest: regression
Number of trees: 500
No. of variables tried at each split: 2
Mean of squared residuals: 8.130213
% Var explained: 86.53
> importance(rf) # Measures reduction of the node’s impurity (diversity), if split by the given X-variable
IncNodePurity
cylinders 2142.2611
displacement 3135.1382
horsepower 1752.5234
weight 2399.9452
acceleration 524.3148
year 1196.4857
origin 546.0370
> varImpPlot(rf)
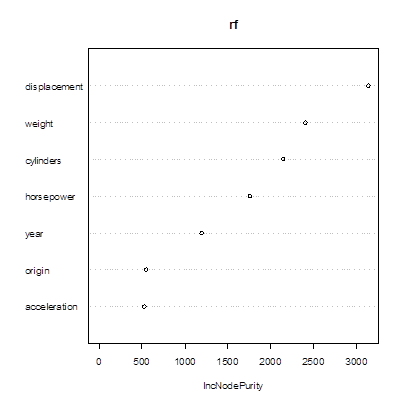
2. Cross-validation
> Z = sample(n,200)
> rf = randomForest(mpg ~ .-name, data=Auto, subset=Z)
> Yhat = predict(rf, newdata=Auto[-Z,])
> mean((Yhat - mpg[-Z])^2)
[1] 10.19022
# The mean-square error of prediction, estimated by the validation set cross-validation, is 10.19022.
3. Searching for the optimal solution
> dim(Auto)
[1] 392 9
# There are 9 variables overall in the data set, minus mpg and name = 7 variables. Let’s sample m = root of 7, rounded = 3.
> rf3 = randomForest(mpg ~ .-name, data=Auto, mtry=3) # mtry is m, the number of X-variables available at each node
> plot(rf3)
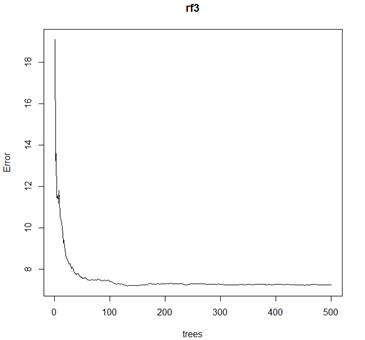
# How many trees to grow? The default is 500, but error is rather flat after 100.
# Random forest tool has multiple output:
> names(rf3)
[1] "call" "type" "predicted" "mse"
[5] "rsq" "oob.times" "importance" "importanceSD"
[9] "localImportance" "proximity" "ntree" "mtry"
[13] "forest" "coefs" "y" "test"
[17] "inbag" "terms"
# We would like to minimize the mean squared error and to maximize R2, the percent of total variation explained by the forest.
> which.min(rf3$mse)
[1] 147
> which.max(rf3$rsq)
[1] 147
# Alright, let’s use 147 trees whose results will get averaged in this random forest.
> rf3.147 = randomForest(mpg ~ .-name, data=Auto, mtry=3, ntree=147)
> rf3.147
Call:
randomForest(formula = mpg ~ . - name, data = Auto, mtry = 3, ntree = 147)
Type of random forest: regression
Number of trees: 147
No. of variables tried at each split: 3
Mean of squared residuals: 7.322948
% Var explained: 87.63
# This is an improvement in both MSE and R2, comparing with our first random forest.
# We can optimize both m and number of trees, by cross-validation.
> Z = sample(n,n-50) # I’m choosing a small test set to make the training set close to the whole data set
> cv.err = rep(0,7) # The optimal random forest may be too dependent on the sample size
> n.trees = rep(0,7)
> for (m in 1:7){
+ rf.m = randomForest( mpg ~ .-name, data=Auto[Z,], mtry=m )
+ opt.trees = which.min(rf.m$mse)
+ rf.m = randomForest( mpg ~ .-name, data=Auto[Z,], mtry=m, ntree=opt.trees )
+ Yhat = predict( rf.m, newdata=Auto[-Z,] )
+ mse = mean( (Yhat - mpg[-Z])^2 )
+ cv.err[m] = mse
+ n.trees[m] = opt.trees
+ }
> which.min(cv.err)
[1] 7
# 7? Apparently, bagging (m=p=7) was the best choice among random forests.
> plot(cv.err); lines(cv.err)
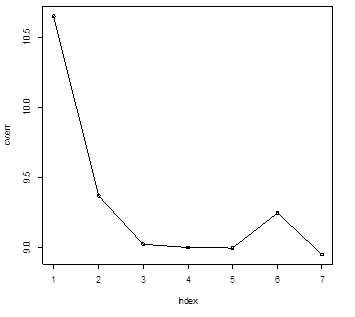
> cv.err
[1] 10.652190 9.368370 9.023726 9.000002 8.996304 9.248892 8.951198
> n.trees
[1] 112 494 318 208 484 319 293
# Result: here is the optimal random forest, which happened to reduce to bagging.
> rf.optimal = randomForest( mpg ~ .-name, data=Auto, mtry=7, ntree=293 )
> rf.optimal
Type of random forest: regression
Number of trees: 293
No. of variables tried at each split: 7
Mean of squared residuals: 7.407668
% Var explained: 87.81
> importance(rf.optimal)
IncNodePurity
cylinders 4820.4219
displacement 7672.7643
horsepower 2838.2213
weight 4603.0577
acceleration 647.7559
year 2861.6324
origin 133.9062